The Scancer Project
Scancer aims to increase access to the early detection of breast cancer.
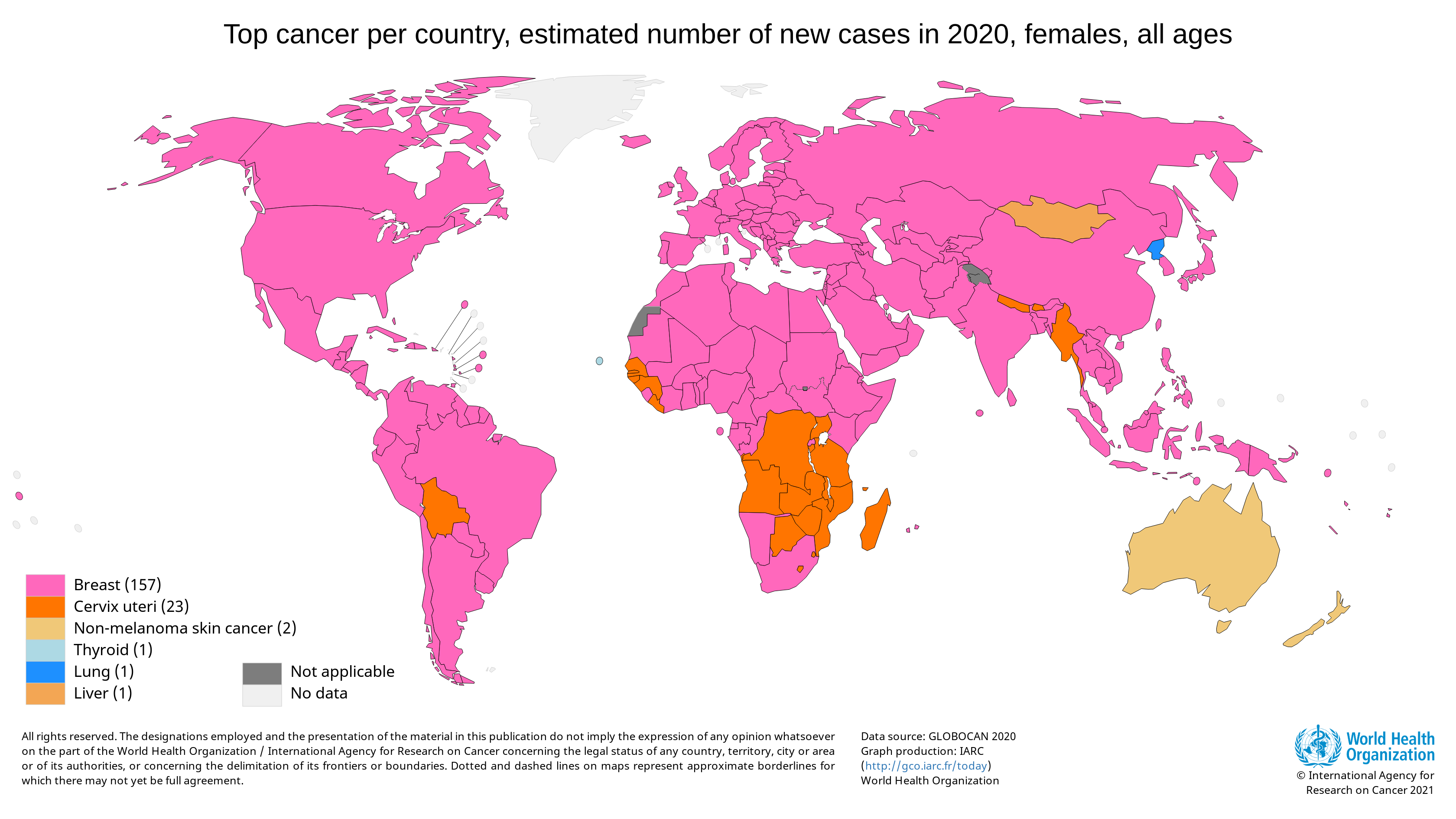
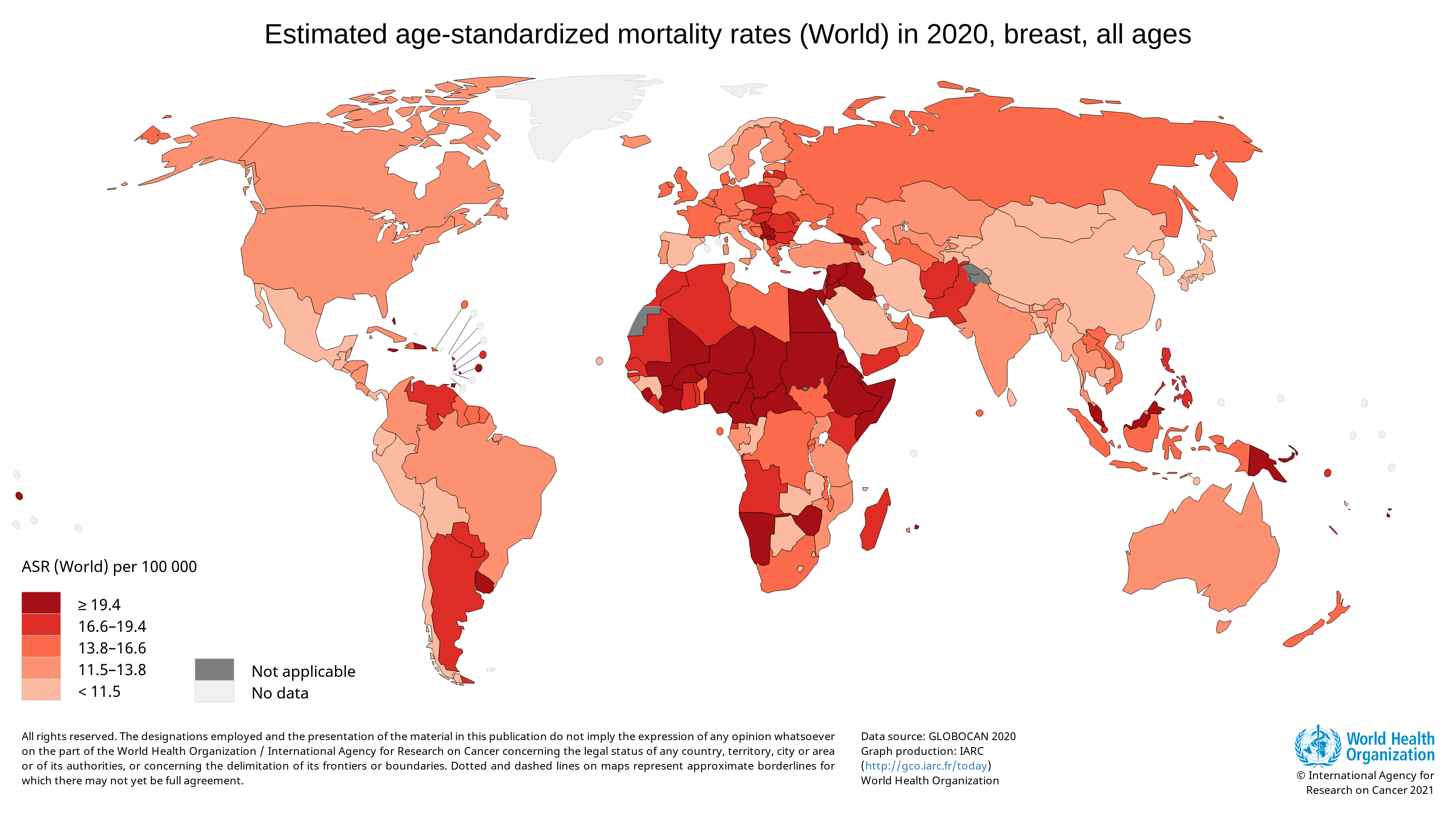
Breast cancer is the most diagnosed cancer among women worldwide. In 2020, there were an estimated 684,996 deaths from breast cancer. A disproportionate number of these deaths occurred in low-resource settings. [ref]
Survival rates for breast cancer are high when the cancer is detected early and where treatment is available. Unfortunately, 50 to 80% of breast cancer cases are diagnosed at an advanced stage in many low- and middle-income countries. By this point, the cancer is usually incurable. [ref]
One of the primary reasons for this is that the tests and procedures used to diagnose this disease (such as mammography and lymph node biopsy) require a lot of human expertise, which is very limited. Fortunately, most of these diagnostic procedures involve visual inspection of scans, which is ideally suited to the strengths of deep learning-based computer vision techniques.
There are many groups that realise this, and so there exist numerous published papers that describe how deep learning models could help improve different aspects of diagnosis. But there isn’t a central place (that we know of) that collects the best of these ideas into a comprehensive solution that is clinically relevant and accessible.
This is exactly what the Scancer project aims to do.
How Do We Propose to Do This?
We have surveyed the specific procedures used to diagnose breast cancer. The graphic that follows depicts these procedures, and highlights in purple the ones that that would most benefit from computer vision.

With this as the background, our proposed solution consists of two major parts:
- First, we maintain a catalogue of the best deep learning vision models for these different parts of the diagnosis chain. These models will be pre-trained, optimised and stored in a model store (TorchServe).
-
Then, on top of this, we construct a software stack that makes these models accessible. The stack we are working on includes:
- An API (powered by TorchServe) that serves these state-of-the-art computer vision models relevant to varying stages of breast cancer detection.
- A web app (written in Django) that wraps this API and sits within a clinically-relevant setting, assisting human experts.
This entire pipeline has been visualised in the following system-level diagram.

Since this is a really open-ended and long term effort, let’s take a closer look at what we have working so far.
What Do We Have Working so Far?
We decided to narrow our focus on just one diagnostic procedure and work out an end-to-end pipeline aound it. The procedure we chose is the analysis of lymph node biopsy scans. During a biopsy, a doctor uses a specialised needle to extract a sample of tissue from the suspicious area. These samples are sent to a lab to be analysed by a pathologist.
The reason we chose this procedure to focus on is:
- A biopsy is the only definitive way to diagnose breast cancer. [ref]
- Manual pathological review of these samples are time-consuming and laborious. [ref].
- Deep learning techniques performing significantly better than human expert baselines. [ref]
- There exists high quality data to train from, e.g. the Camelyon 16 Grand Challenge.
Because the data from the Camelyon 16 challenge is really large (of the order of 2 GB per slide scan), our initial training efforts have focused on a simplified dataset derived from it. This is called the PatchCamelyon (PCam) dataset, and consists of 327,680 color images (96px × 96px) extracted from the full histopathologic scans of lymph node sections. We then proceeded to perform inference on some samples from the larger original data.
Our progress thus far is documented in the table below.
| # | Component | Technologies | What Can You Learn? | GitHub |
|---|---|---|---|---|
| 1 | Trained classification model for the Patch Cameleyon (PCam) dataset | PyTorch, Google Colab, H5py, Weights & Biasis | How to train your own cancer detection model. | Repo |
| 2 | Scancer Model Store | TorchServe | How to package your detection model for serving, and how to contribute it to the Scancer model store. | Docs |
| 3 | Scancer API | TorchServe | How to use the Scancer REST API as part of our primary web app or in your own app. | Docs |
| 4 | Scancer Web App | Django, Django REST Framework, Postgresql, Celery, Redis, Gunicorn, Nginx | How to structure a Django web app for production use. | Repo |
| 5 | Offline inference to generate a heatmap on samples of the Cameleyon 16 dataset | PyTorch, Google Colab, OpenSlide | How to slide a model trained on patches to perform inference on a whole slide image. | Notebook |
| 6 | Scancer Setup Scripts | Ansible | How to setup your own production-ready Django web app wrapping a TorchServe API using reproducible scripts. | Repo |
For a more visual indication of progress, here are some screenshots of the different components. The screenshots cover the functioning pipeline, as well as a web app prototype that aims to gather feedback from domain experts.
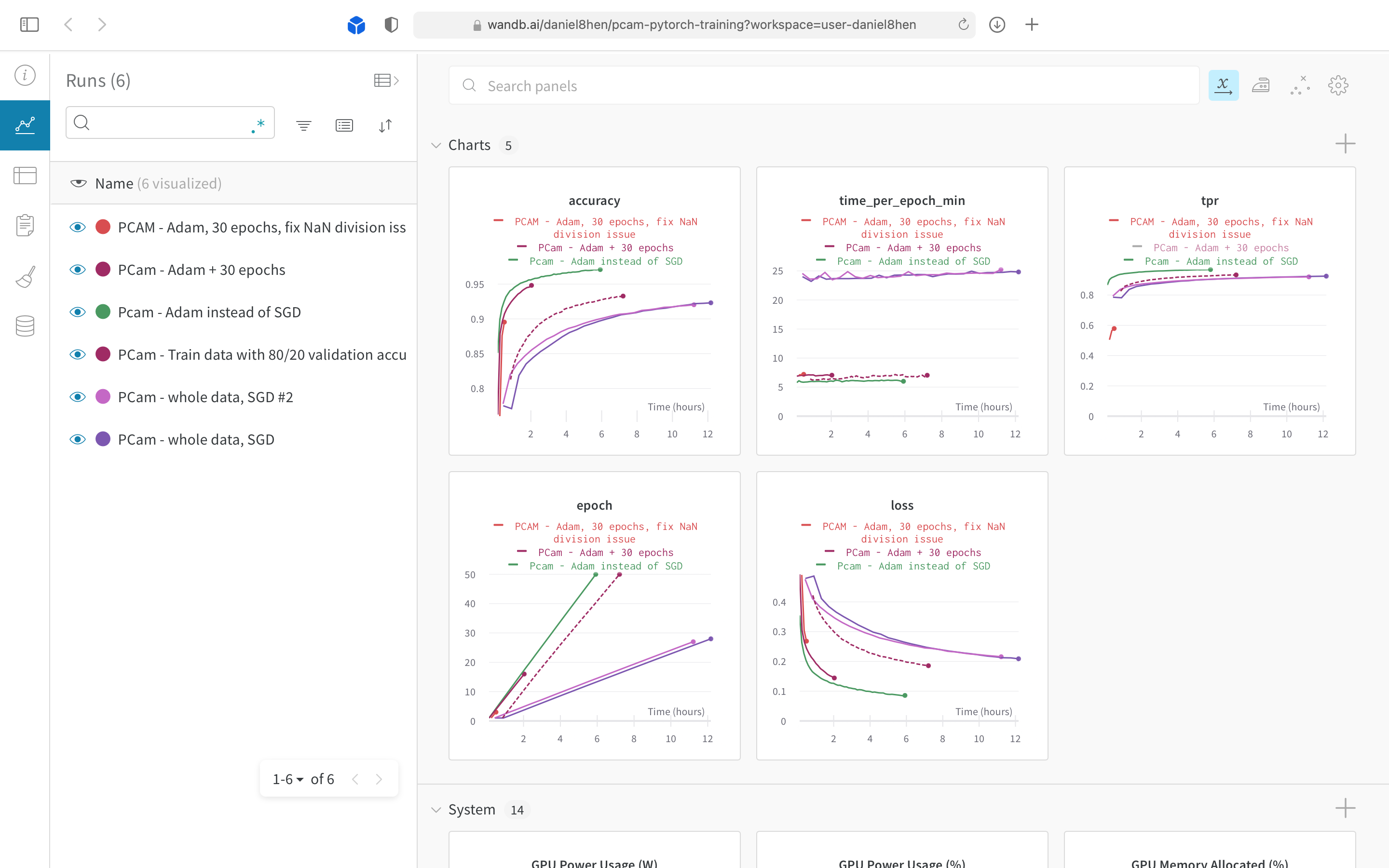
Tuning hyperparameters for our PCam CNN model.
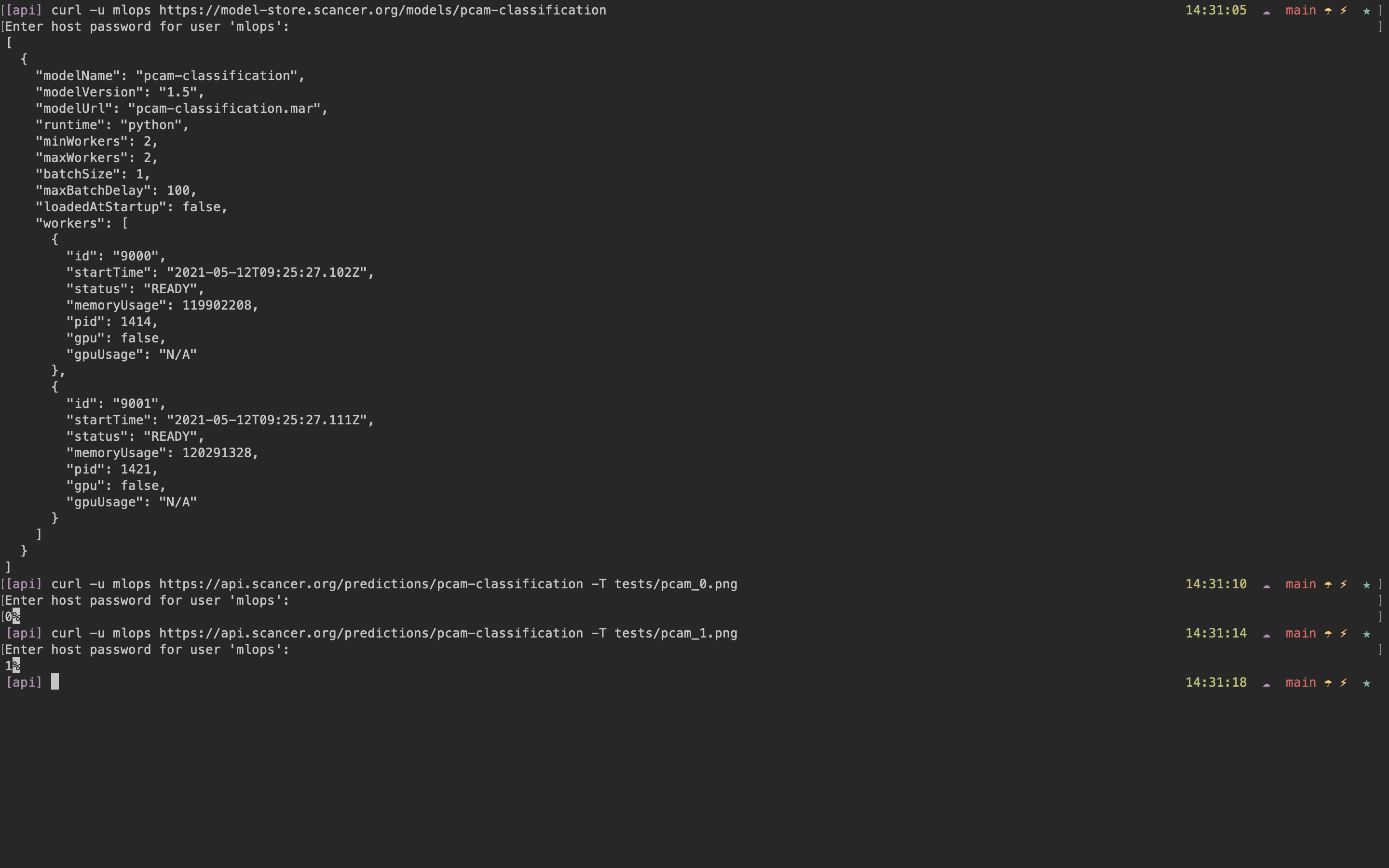
Seeing the model in the model store, and using it as an API.
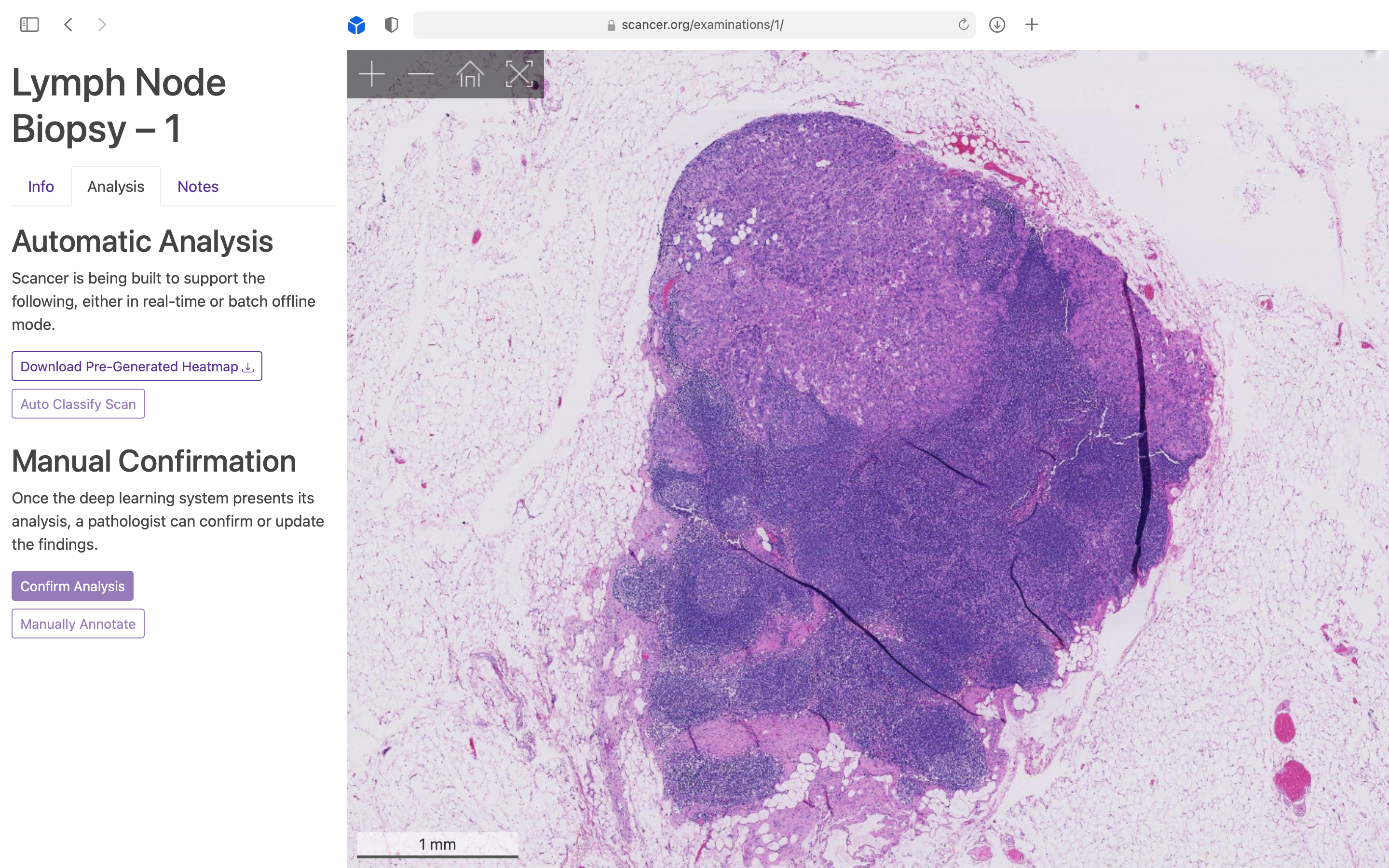
Visualisation of a Whole Slide Image (WSI) using a Google Maps style zoom interface.
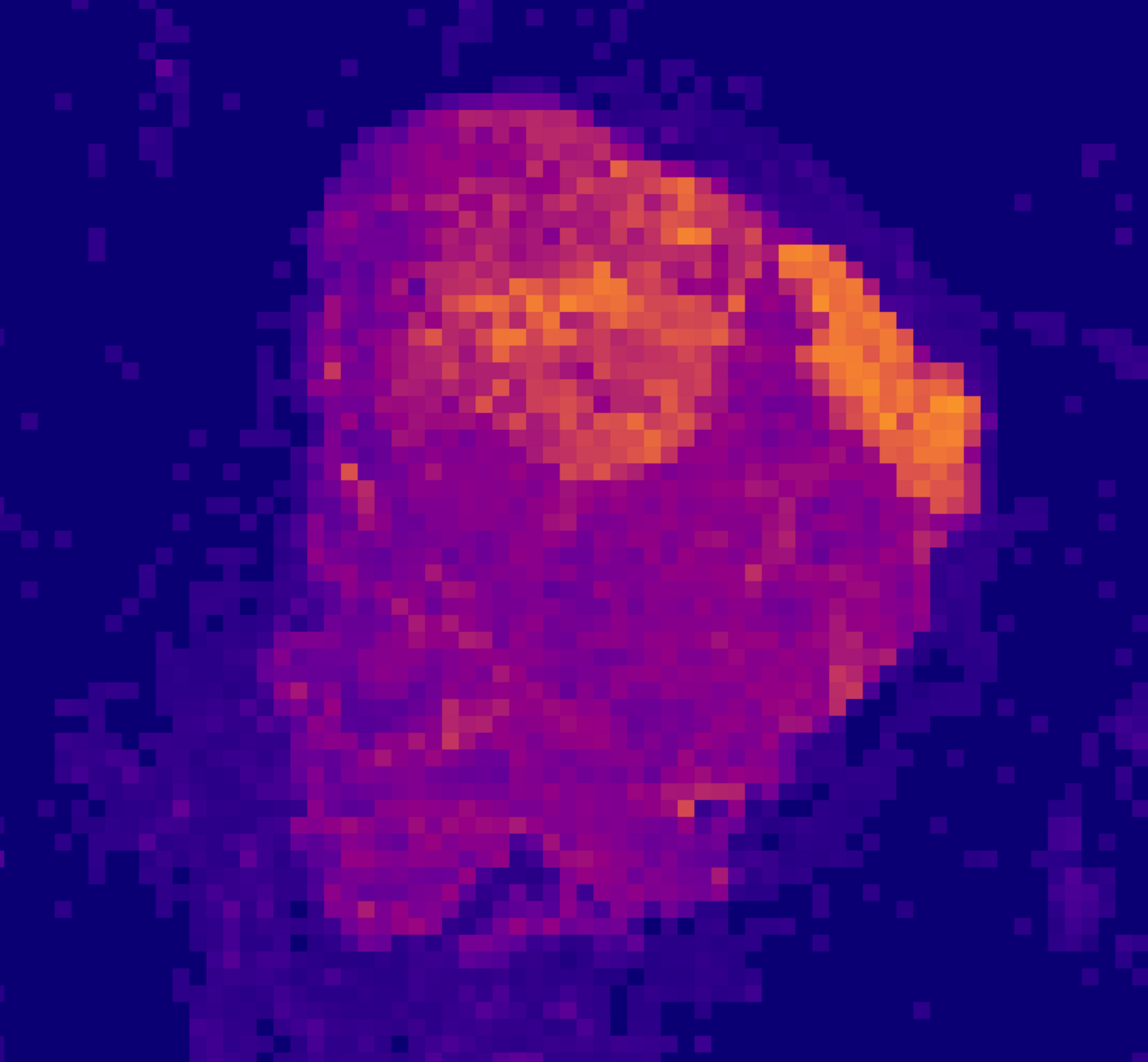

An offline rendered heatmap to highlight potential regions with cancerous tissue.
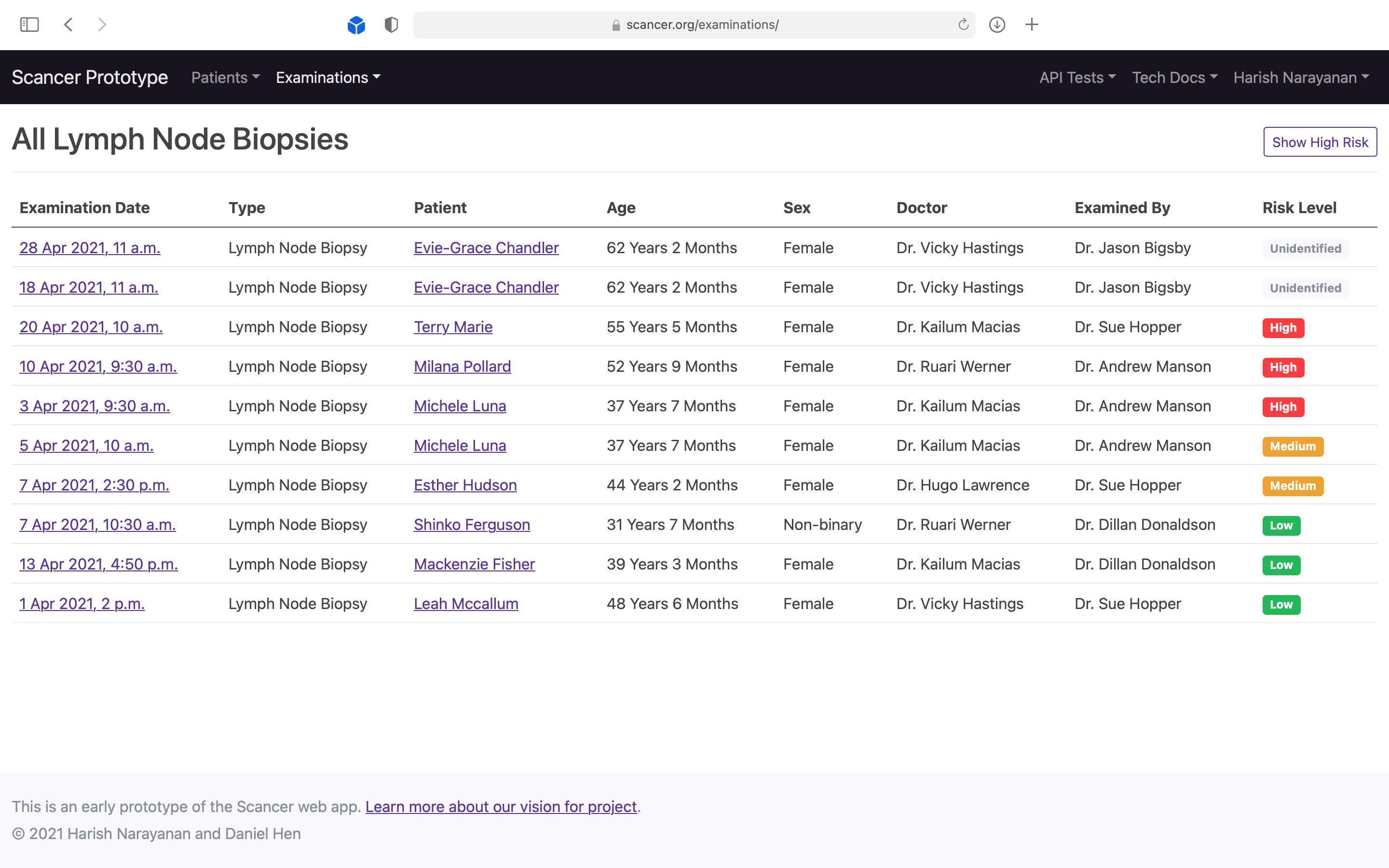
Demonstrating to pathologists that the system could be used to auto-prioritise incoming scans by risk level.
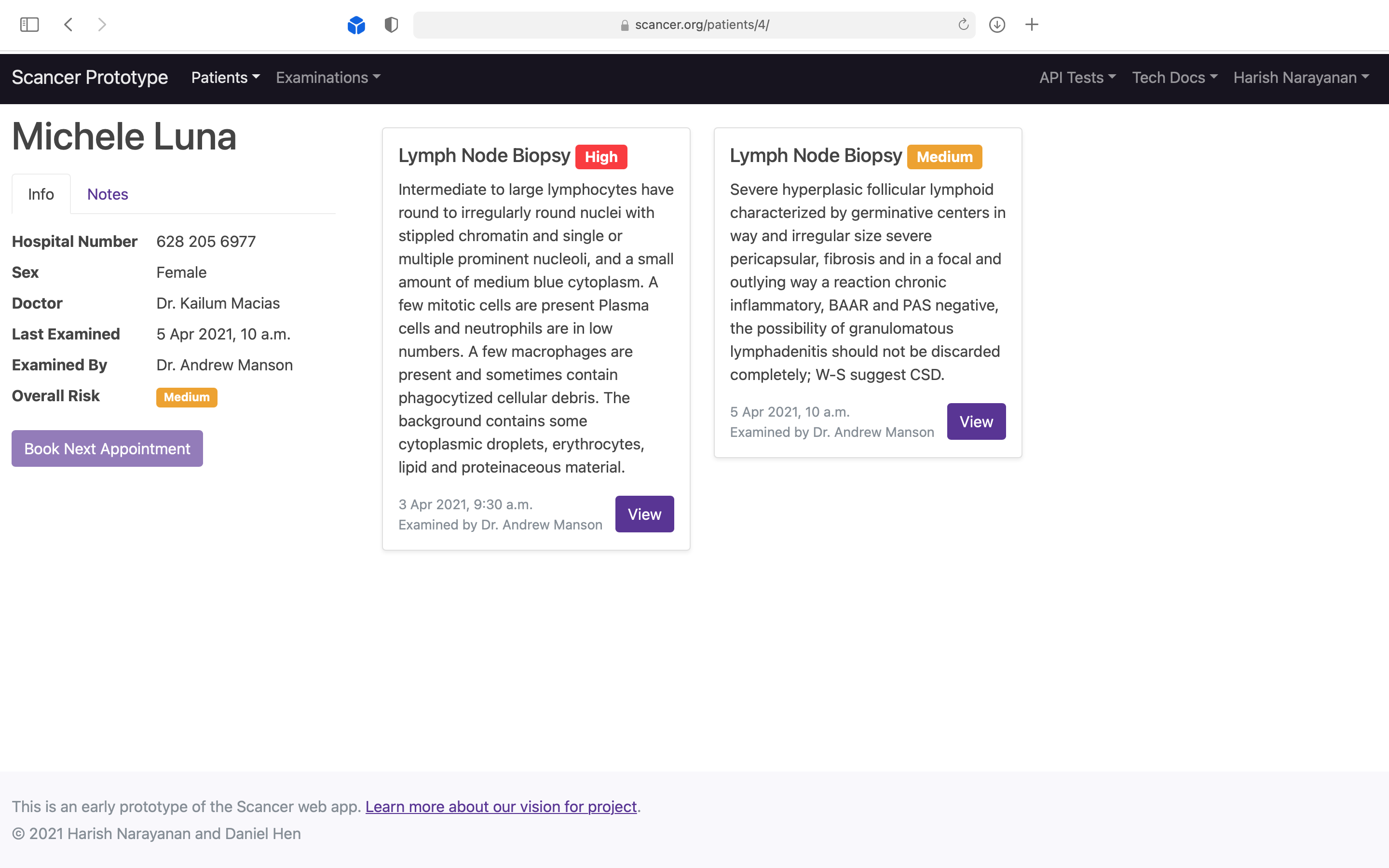
Demonstrating to doctors that they can see all scan results in one view, allowing them to decide the next step.
Where We Are Going Next, and How You Can Help
The project we have embarked on is really open-ended and long term. Thus far, we have a functioning ML pipeline, an example model, and a prototype web app that sits on top of it. We will continue to extend this app and underlying model until it is genuinely useful for histopathologists.
Everything we’ve built is released open source (MIT License) and in a reproducible form, so that anyone can learn from, use, and contribute to the effort. Going forward, we will encourage the community to contribute models and workflows for other diagnostic procedures.
In particular:
- Domain experts can help with labelling data, provide their own unique datasets, and help us understand issues in their everyday work
- Machine learning practitioners can help train even more robust models
- Companies can help with compute budgets, etc.
If you are interested in helping or finding out more, please Sign Up or Write to Us